DADPS H2/D2 Biotin Azide Pack
DADPS H2/D2 Biotin Azide probes enable mass-independent chemical proteomics platform that helps to address unique challenges of the proteome characterization. In this approach a unique isotopic signature is embedded exclusively into the peptides and it serves as a computationally recognizable full-scan MS reporter. A computational algorithm, termed isotopic signature transfer and mass pattern prediction (IsoStamp), for the detection of recoded species in full-scan mass spectra, was also developed by the Carolyn Bertozzi group. DADPS H2/D2 Biotin Azide pack contains 2 mg of each probe.
While there has been much interest in profiling the intact glycoproteome, the complexity of glycoproteoforms (and more broadly, all proteoforms) remains challenging to completely define. Mass spectrometry (MS) is commonly employed for characterization of complex proteomic samples. A popular strategy for protein identification is the bottom-up shotgun proteomics approach. In this method, a mixture of proteins is subjected to proteolytic digestion, the resulting peptides are separated by LC and detected by MS, and their parent proteins are inferred from the assigned peptide sequences.
To convert MS data acquired from proteolytic digests into protein identifications, tandem MS can be used to obtain sequence information for individual peptides, followed by comparing an in-silico proteolytic digest of an organism’s proteome. Typically, only the most abundant peptides are selected for fragmentation (Figure 2), whereas data for those peptides in relatively low quantities are not obtained. An inherent problem in shotgun proteomics is identifying proteins of low abundance, such as biomarkers for disease states, against a background of proteins whose concentrations can span up to 12 orders of magnitude.
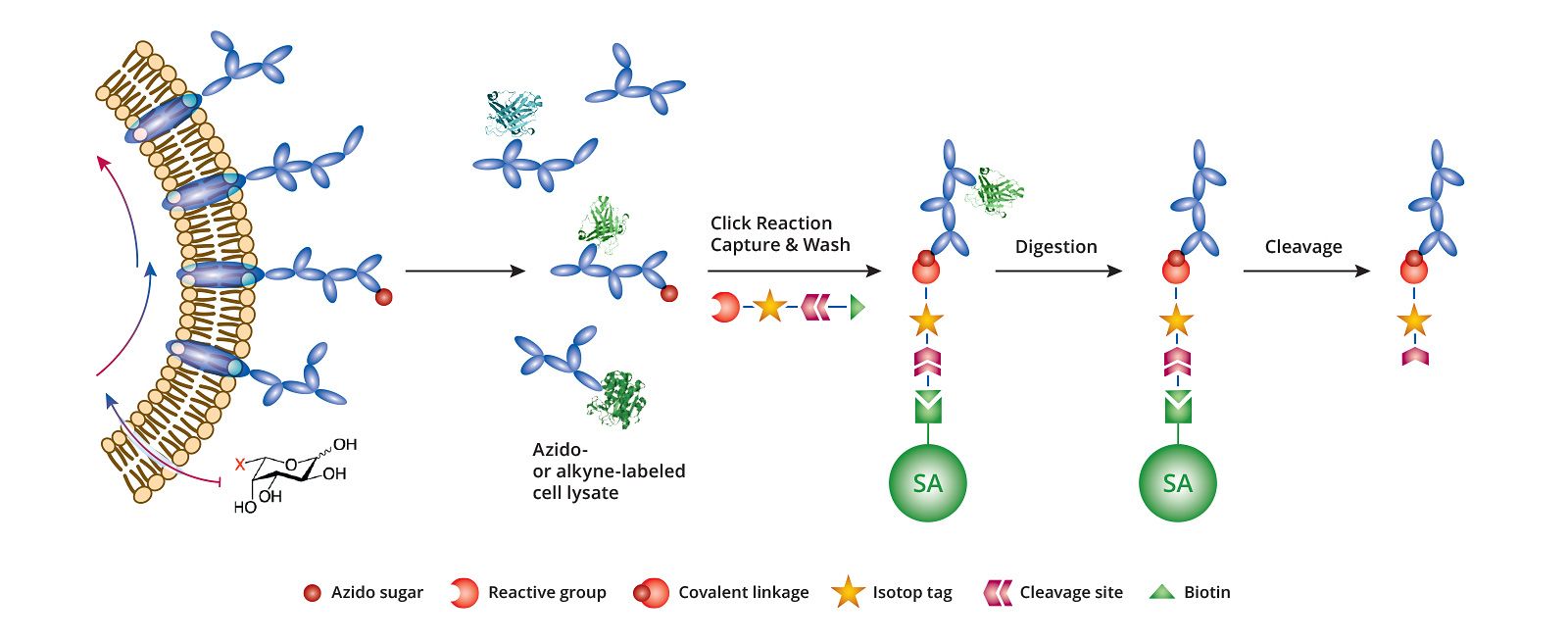 Figure 1. Metabolic labeling with a chemically functionalized glycan, chemical taggingand enrichment using an isotopic recoding affinity probe
Figure 1. Metabolic labeling with a chemically functionalized glycan, chemical taggingand enrichment using an isotopic recoding affinity probe
To address the unique challenges of the global characterization of the intact glycoproteome, a mass-independent chemical glycoproteomics platform, termed isotope targeted glycoproteomics (IsoTag) was developed by the Carolyn Bertozzi group. The platform is comprised of four central components: (i) metabolic labeling with a chemically functionalized glycan, (ii) chemical tagging and enrichment using an isotopic recoding affinity probe, (iii) directed tandem MS, and (iv) targeted glycopeptide assignment (Figure 2).
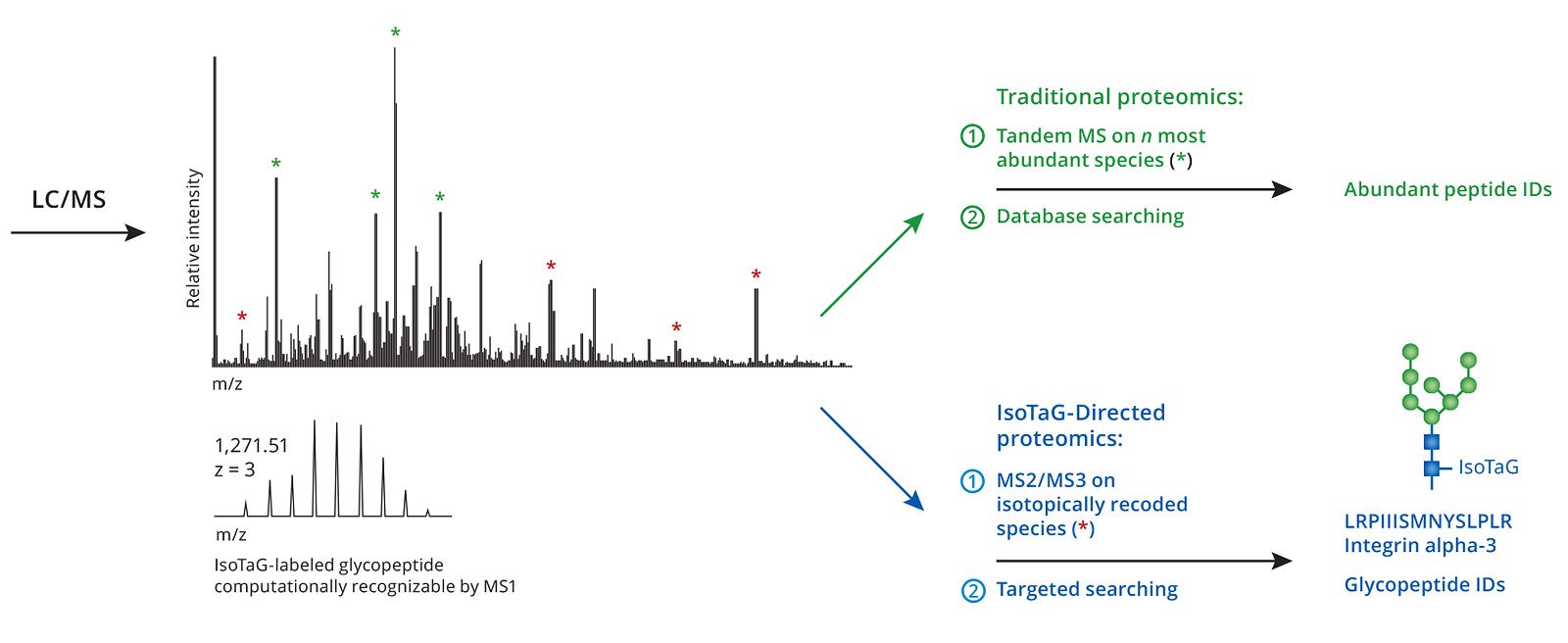
Figure 2. Traditional proteomics and Iso-Tag-directed proteomics workflow
IsoTaG is performed by isotopic recoding and enrichment of metabolically labeled glycoproteins followed by directed tandem MS (MS2 or MSn) analysis and intact glycopeptide assignment. Isotopic recoding is accomplished by metabolic labeling of cell or tissue samples with azide- or alkyne-functionalized sugars, followed by chemical conjugation with a biotin probe bearing a unique isotopic signature.
Some examples of sugar labels are peracetylated N-azidoacetylmannosamine (Ac4ManNAz), which is converted to the corresponding azidosialic acid (SiaNAz), and peracetylated N-azidoacetylgalactosamine (Ac4GalNAz), which is metabolized to label glycans possessing N-acetylglucosamine (GlcNAc) or N-acetylgalactosamine (GalNAc) (not provided with kit).
In order to perform isotopic tagging, the kit provides two cleavable IsoTaG probes encoded by zero [M] and two [M + 2] deuterium atoms. Probes with different encoding can be provided by Click Chemistry Tools though custom synthesis. The IsoTaG probe with zero, and that with two deuterium atoms [M, M + 2] can be used in different proportions; 1:1, 1:2, 1:3 and 1:4. Pattern recognition with isotopic ratio of 1:3 showed the highest fidelity.
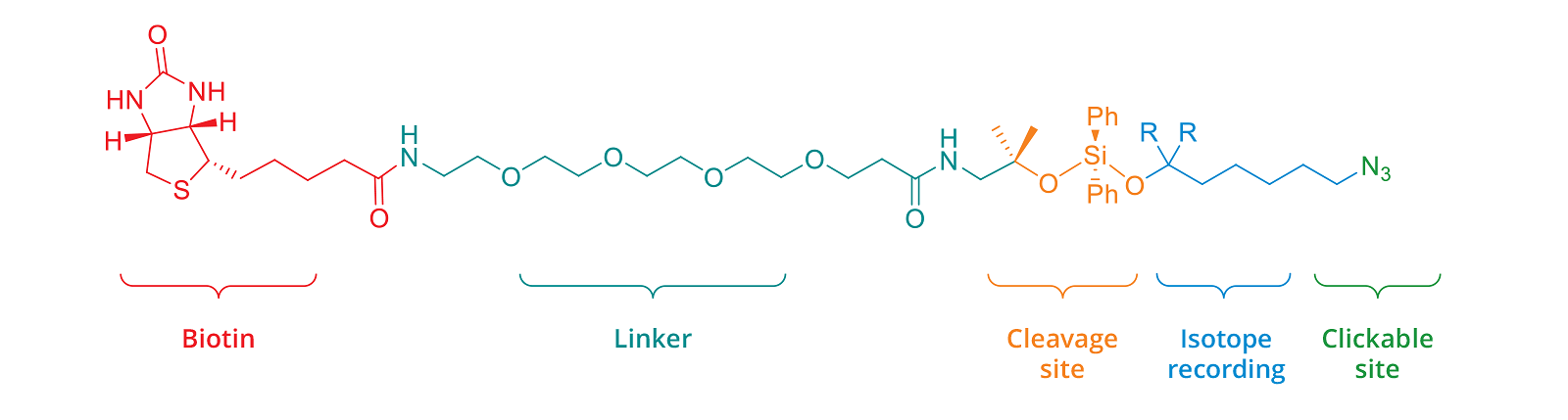
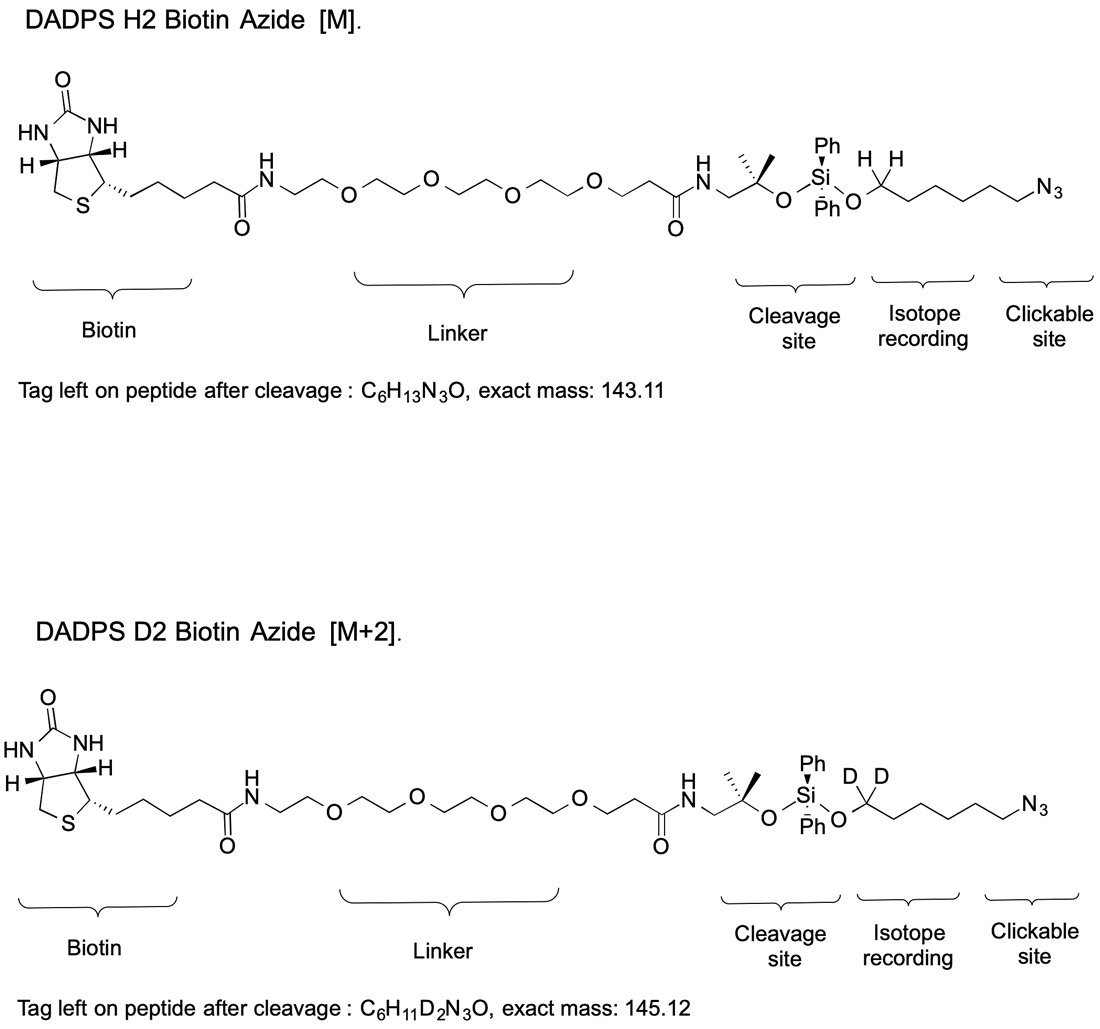
Figure 3. Cleavable IsoTaG probe encoded by zero deuterium atoms [M] (R = H) and two deuterium atoms [M+2] (R = D)
Through these probes, a unique isotopic signature is embedded exclusively into the glycopeptides. The isotopic signature serves as a computationally recognizable full-scan MS reporter. A computational algorithm, termed isotopic signature transfer and mass pattern prediction (IsoStamp), for the detection of recorded species in full-scan mass spectra, was also developed by the Carolyn Bertozzi group. IsoStamp compares observed and predicted isotopic envelopes to identify chemically tagged species in full-scan mass spectra.
IsoTag has the potential to enhance any proteomics platform that employs chemical labeling for targeted protein identification, including isotope-coded affinity tagging, isobaric tagging for relative and absolute quantitation, and chemical tagging strategies for post-translational modification.
Selected References:
1. Woo, C. M., et al. (2017). Development of IsoTaG, a Chemical Glycoproteomics Technique for Profiling Intact N- and O?Glycopeptides from Whole Cell Proteomess. J. Proteome Res., 16: 1706?18.
2. Woo, C.M.., et al. (2017). Mapping and Quantification of Over 2000 O-linked Glycopeptides in Activated Human T Cells with Isotope-Targeted Glycoproteomics (Isotag). Mol. Cell.Proteomics., 17: 764?75.
3. Gao, G., et al. (2017). Small Molecule Interactome Mapping by Photoaffinity Labeling Reveals Binding Site Hotspots for the NSAIDs. J. Am. Chem. Soc., 140: 4259?68.
4. Woo, C.M., et al. (2015). Isotope-targeted glycoproteomics (IsoTaG): a mass-independent platform for intact N- and O-glycopeptide discovery and analysis. Nat Methods., 12: 561?7.
5. Weerapana, E., et al. (2010). Quantitative reactivity profiling predicts functional cysteines in proteomes. Nature, 648: 790?5.
Iso-Tag products are covered by U.S. Patent No.: 10,114,026.
This product may be used for research purposes only. It is not licensed for resale and may only be used by the buyer. This product may not be used and is not licensed for clinical assays, where the results of such assays are provided as a diagnostic service. If a diagnostic or therapeutic use is anticipated, then a license must be requested from the University of California. The availability of such diagnostic and therapeutic use license(s) cannot be guaranteed from the University of California.
1. Woo, C.M., et al. (2017). Development of IsoTaG, a Chemical Glycoproteomics Technique for Profiling Intact N- and O?Glycopeptides from Whole Cell Proteomess. J. Proteome Res., 16: 1706?18. [PubMed]
2. Woo, C.M., et al. (2017). Mapping and Quantification of Over 2000 O-linked Glycopeptides in Activated Human T Cells with Isotope-Targeted Glycoproteomics (Isotag). Mol. Cell. Proteomics 17: 764-75. [PubMed]
3. Goa, G., et al. (2017). Small Molecule Interactome Mapping by Photoaffinity Labeling Reveals Binding Site Hotspots for the NSAIDs. J. Am. Chem. Soc., 140: 4259?68. [PubMed]
4. Woo, C.M., et al. (2015). Isotope-targeted glycoproteomics (IsoTaG): a mass-independent platform for intact N- and O-glycopeptide discovery and analysis.Nat Methods., 12: 561-7. [PubMed]
5. Weerapana, E., et al. (2010). Quantitative reactivity profiling predicts functional cysteines in proteomes. Nature 468: 790?5. [PubMed]
Abs/Em Maxima: 346/445 nmExtinction Coefficient: 19,000Spectrally Similar Dyes: Alexa Fluor? 350, CF? 350, DyLight 350, AMCAAZDye? 350 is a blue-fluorescent azide-activated probe that reacts with terminal alkynes via a copper-catalyzed click react…
Abs/Em Maxima: 402/424 nmExtinction Coefficient: 35,000Flow Cytometry Laser Line: 405 nmMicroscopy Laser Line: 405 nmSpectrally Similar Dyes: Alexa Fluor? 405, CF? 405, Cascade Blue?, DyLight? 405AZDye? 405 Azide is a water-soluble, pH-insensiti…
Abs/Em Maxima: 346/445 nmExtinction Coefficient: 19,000Spectrally Similar Dyes: Alexa Fluor? 350, CF? 350, DyLight 350, AMCAAZDye? 350 Alkyne (Alexa Fluor? 350 Alkyne equivalent) is a blue-fluorescent, alkyne-activated probe that reacts with azid…
Abs/Em Maxima: 430/537 nmExtinction Coefficient: 15,000Spectrally Similar Dyes: Alexa Fluor? 430, CF? 430AZDye? 430 Azide is a water-soluble, green-fluorescent azide-activated probe that reacts with terminal alkynes via a copper-catalyzed click re…
Abs/Em Maxima: 346/445 nmExtinction Coefficient: 19,000Spectrally Similar Dyes: Alexa Fluor? 350, CF? 350, DyLight 350, AMCAAZDye? 350 DBCO reacts with azides via a copper-free “click chemistry” reaction to form a stable triazole and does not re…
Abs/Em Maxima: 402/424 nmExtinction Coefficient: 35,000Flow Cytometry Laser Line: 405 nmMicroscopy Laser Line: 405 nmSpectrally Similar Dyes: Alexa Fluor? 405, CF? 405, Cascade Blue?, DyLight? 405AZDye? 405 Alkyne reacts with azides via a copper…
Abs/Em Maxima: 494/517 nmExtinction Coefficient: 73.000Flow Cytometry Laser Line: 488 nmMicroscopy Laser Line: 488 nmSpectrally Similar Dyes: Fluorescein, Alexa Fluor? 488, CF? 488A, DyLight? 488, Atto? 488AZDye? 488 Azide (Alexa Fluor? 488 Azi…
Abs/Em Maxima: 402/424 nmExtinction Coefficient: 35,000Flow Cytometry Laser Line: 405 nmMicroscopy Laser Line: 405 nmSpectrally Similar Dyes: Alexa Fluor? 405, CF? 405, Cascade Blue?, DyLight? 405ZDye? 405 DBCO reacts with azides via a copper-fr…
